Work Packages
An integrated framework combining multi‑omics, functional neurobiology, and genome editing to dissect metabolic control of puberty.
Overview
DOPA‑Kiss is organized into three complementary Work Packages (WP1–WP3) to uncover how hypothalamic Kiss1 circuits link metabolism to pubertal control, with emphasis on early‑onset obesity and sex differences. WP1 charts molecular landscapes of hypothalamus and Kiss1 neurons across pubertal maturation and obesity. WP2 defines lipid and nutrient‑sensing mechanisms, including FFAR/PPAR/TGR5 and cellular energy sensors. WP3 performs in vivo genome editing to test causality and translational relevance.
WP1
Multi‑omics characterization of hypothalamic kisspeptin neurons.
Aim:
To profile molecular, transcriptomic, and metabolic states of hypothalamus and Kiss1 populations under normal conditions and early‑onset obesity.
Key approaches:
✅ Comprehensive metabolomic and lipidomic profiling of the hypothalamus.
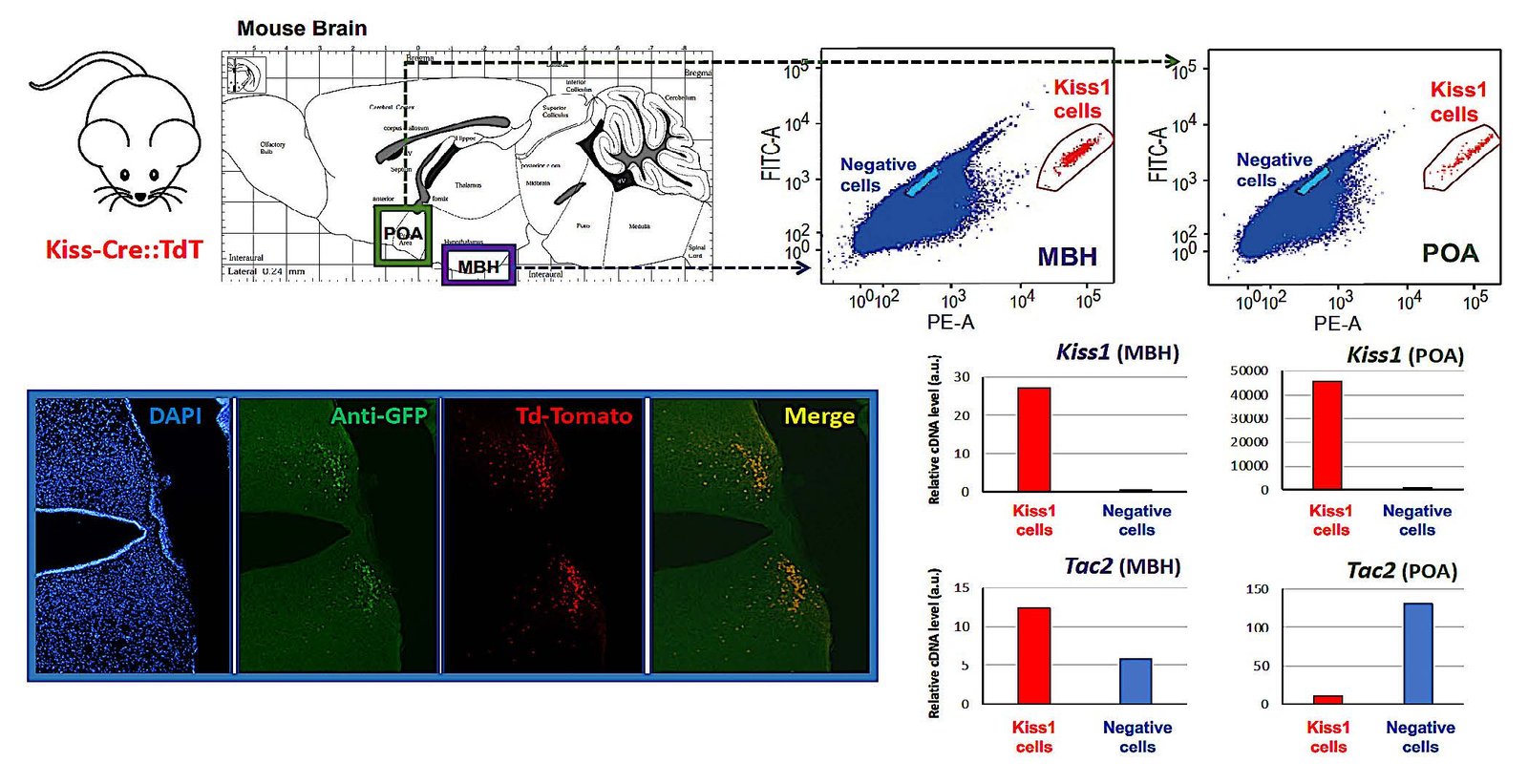
✅ Isolation and profiling of Kiss1 neuronal populations for epigenomic and single‑cell analyses.
✅ Spatial multi‑omics to map in situ molecular alterations across development.
Expected outcome:
An integrated atlas revealing deregulated pathways and candidate targets for functional testing in WP2 and WP3.
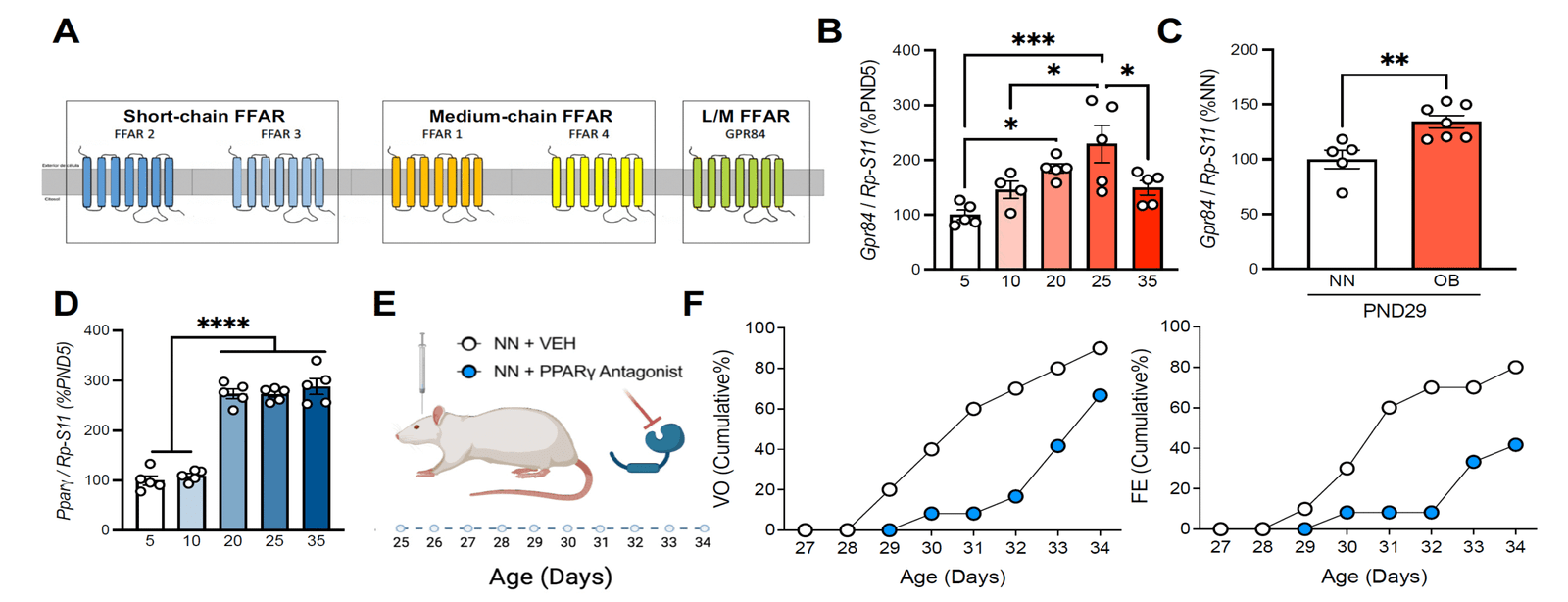
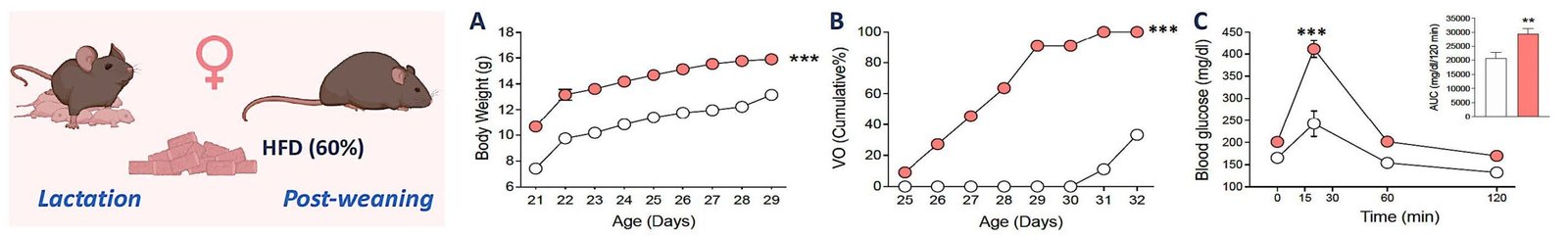
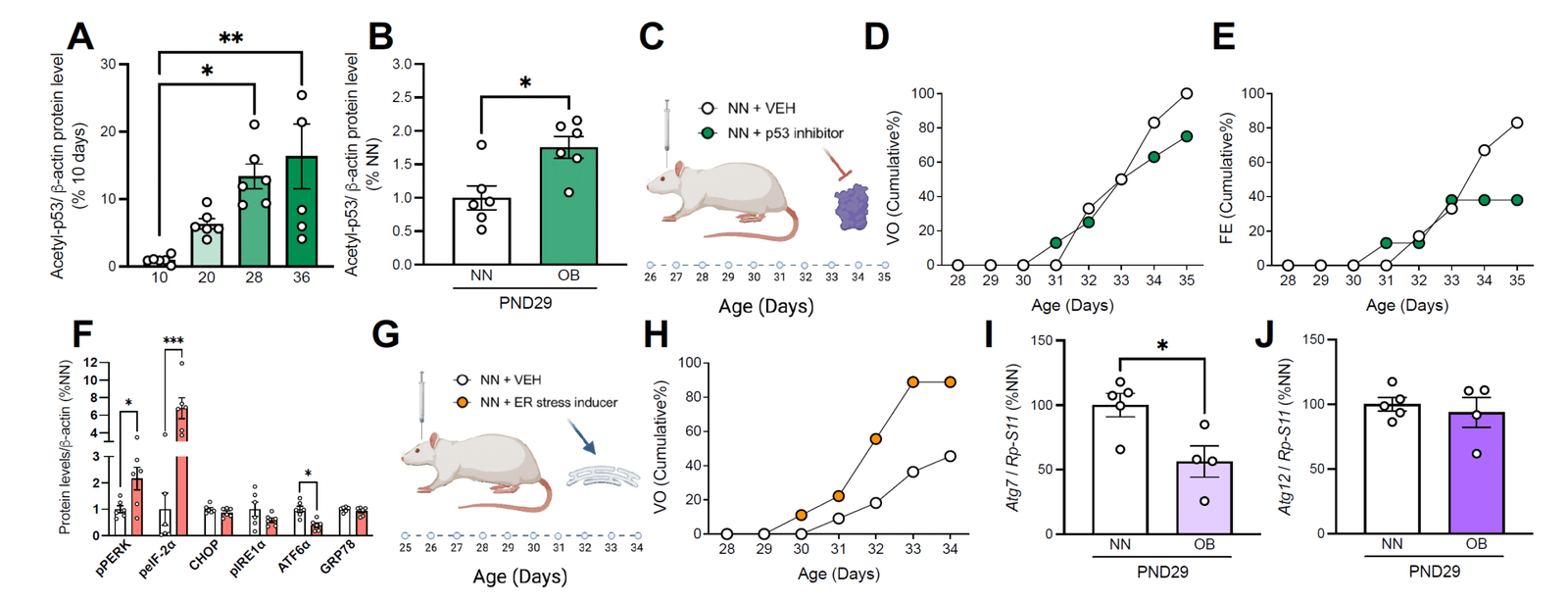
WP2
Mechanistic analysis of metabolic modulation
Aim:
To determine how lipid and nutrient‑driven signals influence hypothalamic circuits and Kiss1 function that regulate pubertal timing.
Key approaches:
✅ Quantitative analyses of gene and protein expression in tissue and sorted neurons.
✅ Genetic and pharmacological interrogation of FFAR/PPAR/TGR5 and energy‑sensing pathways (e.g., AMPK/mTOR/SIRT1).
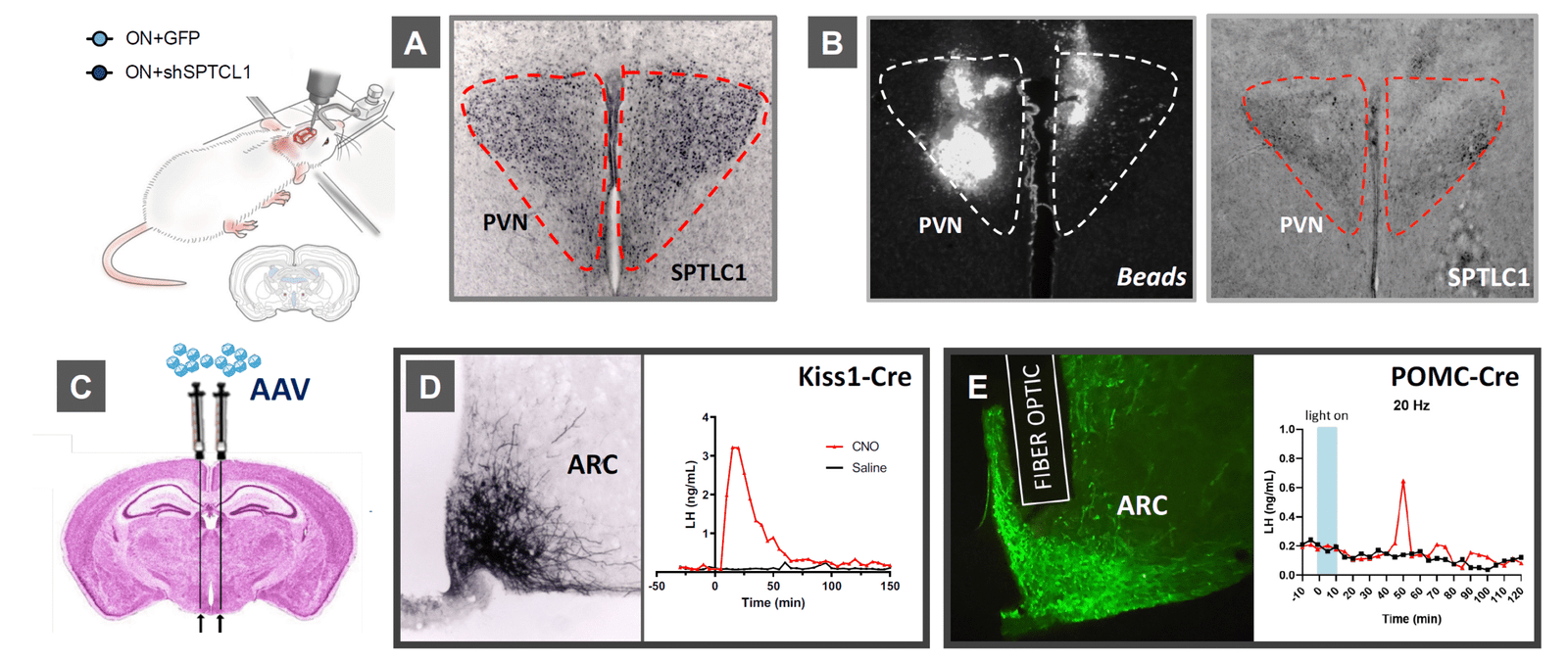
✅ In vivo functional assays in Kiss1 circuits to connect pathway activity with pubertal phenotypes.
Expected outcome:
Identification of circuit‑ and molecule‑level mechanisms by which metabolic cues modulate puberty, including sex‑specific effects.
WP3
Causal validation through targeted genome editing
Aim:
To establish direct causal relationships between prioritized metabolic pathways and pubertal control in Kiss1 neurons.
Key approaches:
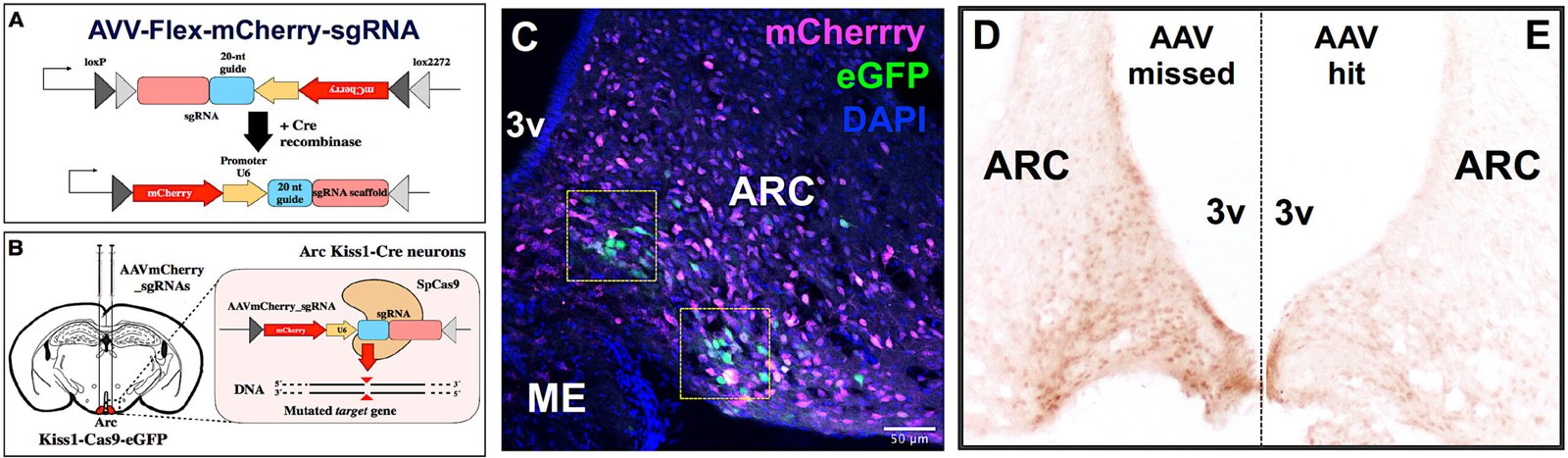
✅ CRISPR/Cas and Cre/LoxP editing targeted to Kiss1 neurons in vivo.
✅ Stereotaxic AAV delivery for spatially resolved manipulation of candidate genes.
✅ Phenotypic analyses of pubertal timing and metabolic endpoints to quantify impact.
Expected outcome:
Definitive causal evidence and validated molecular targets in the central regulation of puberty.